User Interface¶
When reviewing variants you’ll have the following view:
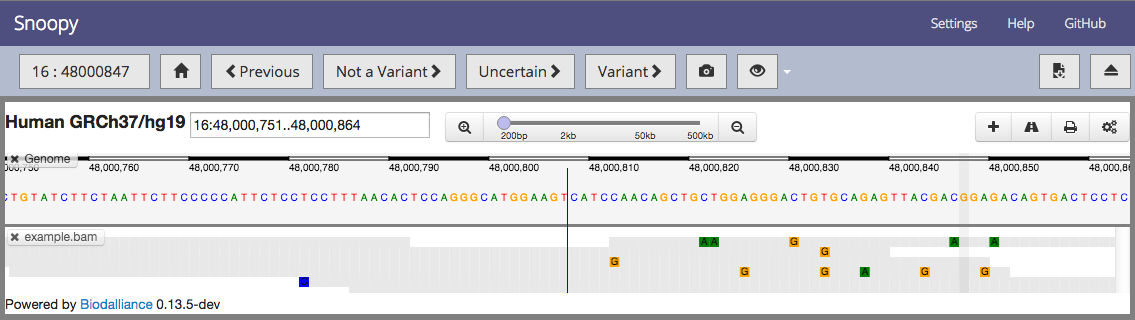
There are two main sections in this view: the Snoopy interface and the Dalliance interface.
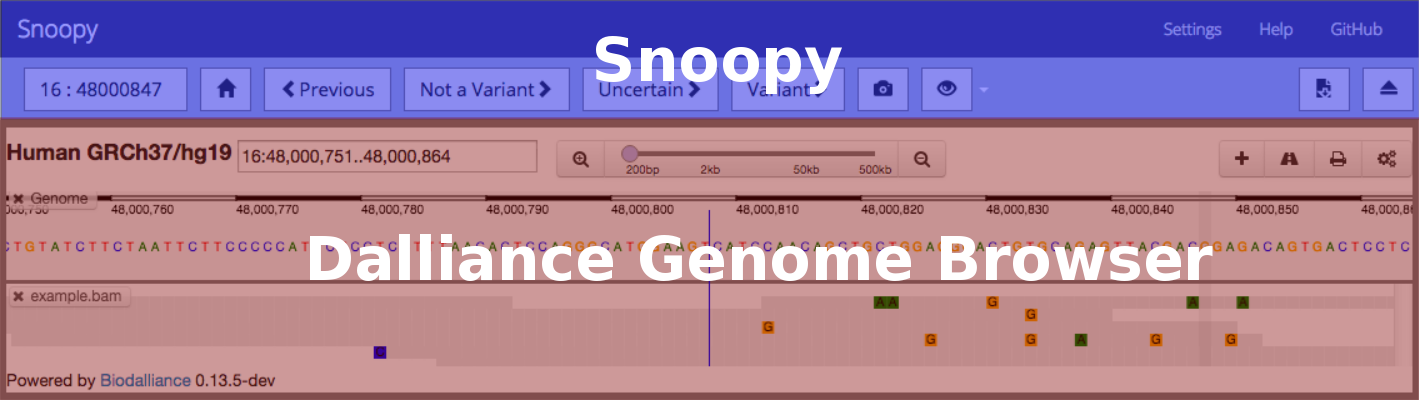
Dalliance¶
Within Dalliance you have access to following:
 - Lists the genome reference being used and the current chromosome and location. You can change the location by entering either a start and end position or a single base position.
- Lists the genome reference being used and the current chromosome and location. You can change the location by entering either a start and end position or a single base position. - Control the zoom depth.
- Control the zoom depth. - Adds a new track to the current view.
- Adds a new track to the current view. - Modify track options such as the max depth, and colors.
- Modify track options such as the max depth, and colors. - Export current view as an SVG or PNG.
- Export current view as an SVG or PNG. - Change some of Dalliance’s options such as vertical guideline location, scrolling direction.
- Change some of Dalliance’s options such as vertical guideline location, scrolling direction.
Snoopy¶
Variant Decisions¶
When either of the decision buttons are clicked, you will advance to the next variant.
 - You are certain that the called site is not actually a variant.
- You are certain that the called site is not actually a variant. - You are unsure if the called site is a variant.
- You are unsure if the called site is a variant. - You are certain that the called variant is truly a variant.
- You are certain that the called variant is truly a variant.
Viewing¶
 - This dropdown button allows you to select four different track styles:
- This dropdown button allows you to select four different track styles:- Raw - Display the bases with color scheme as specified in the settings.
- Condensed - Only display a base if it differs from the reference. If a read exists and is in agreement with the reference, use match color as set in your settings.
- Mismatch - Color code plus strand / minus strand if reference agreement exists. If a base differs, display with the base color as given in settings for raw.
- Coverage - Presents a histogram of coverage. If more than 20% of the bases differ from reference, the proportion of bases are displayed with their bases colors.
 - Clicking this will take a snapshot of whatever view is currently loaded into Dalliance.
- Clicking this will take a snapshot of whatever view is currently loaded into Dalliance.
Admin¶
 - Stop and restart Snoopy button, there will be a prompt asking if you wish to save your progress.
- Stop and restart Snoopy button, there will be a prompt asking if you wish to save your progress. - Save whatever progress you have made so far. The output format is described in Saving Results.
- Save whatever progress you have made so far. The output format is described in Saving Results.
Settings¶
View and change the following settings:
- Connections
- Default Remote, Local and SSH-Bridge settings (URL, credential requirements)
- Dalliance zoom level
- How deep should Dalliance be zoomed into after variant decision
- Color settings
- nucleotide bases colors
- Mismatch style
- Plus/minus strand colors
- Show insertions
- Reflect base quality with transparency
- Condensed style
- Match color
- Reflect base quality with transparency
- Coverage histogram style
- Allele threshold (between 0 and 1): the minimum ratio of allele frequencies before a mismatch is displayed
- Height
- Snapshots
- Automatically take snapshots at each variant
Help¶
A link to the documentation hosted on Read The Docs.
GitHub¶
A link to the GitHub repository holding the snoopy’s source code.
 - This displays the current variant and a QC decision, if available. If you click this button you will presented with a window which summarizes all of the QC decisions made so far, as well as allowing you to quickly navigate to a different variant.
- This displays the current variant and a QC decision, if available. If you click this button you will presented with a window which summarizes all of the QC decisions made so far, as well as allowing you to quickly navigate to a different variant. - Returns to the current variant of interest if you’ve dragged the Dalliance track away.
- Returns to the current variant of interest if you’ve dragged the Dalliance track away. - Go to the previous variant. Clicking this button does not register any variant decisions.
- Go to the previous variant. Clicking this button does not register any variant decisions.